[NiFe] Group 4a-hydrogenase
This entry was last updated at: Feb. 9, 2018, 7:09 a.m.
Properties
| Group | [NiFe] Group 4: Respiratory H2-evolving [NiFe] hydrogenases |
| Subgroup | [NiFe] Group 4a: Formate hydrogenlyases |
| Function | Directly couples formate oxidation to H2 evolution. It remains unclear whether this process can be coupled to proton-translocation through the transmembrane modules. Reverse reaction occurs in vitro but unresolved whether it occurs in vivo. |
| Activity | H2-evolution |
| Oxygen tolerance | O2-sensitive |
| Localisation | Membrane-bound |
Distribution
| Ecosystem distribution | Animal gastrointestinal tracts |
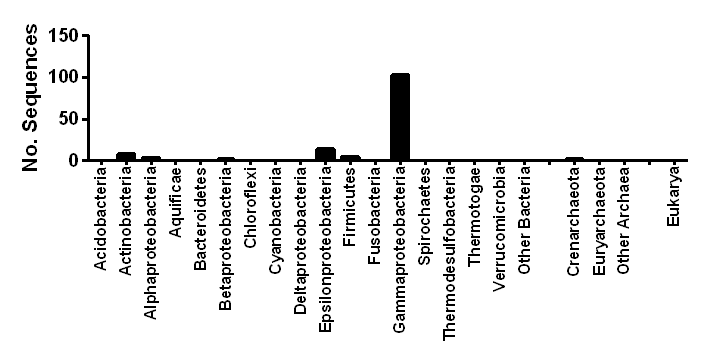
| Taxonomic distribution | Encoded in Enterobacteria and several other Proteobacteria |
Genetic Organisation
Organisation 1: Escherichia coli hyc operon (WP_014639275.1):

Organisation 2: Escherichia coli hyf operon (WP_014641051.1):

Architecture
| Structures |
|
| Subunits | 6 or 9 |
| Subunit description | HycE / HyfG (hydrogenase large subunit) HycG / HyfI (hydrogenase small subunit) HycF / HyfH (iron-sulfur protein) HycB / HyfA (iron-sulfur protein) HycC / HyfB (NuoL-like transmembrane adaptors) HycD / HyfC (NuoH-like transmembrane adaptors) Hyf-type enzymes additionally associated with three antiporter-like subunits (HyfD, HyfE, HyfF) Loosely associated with formate dehydrogenase module (FdhF) |
| Catalytic site | [NiFe]-centre |
| FeS clusters | Proximal: 4Cys[4Fe4S] Medial: Absent Distal: Absent |
Important Notes
It remains to be resolved whether the formate hydrogenlyase is fermentative (proton-nontranslocating) or respiring (proton-translocating) complex. Based on structural analysis of Complex I suggests, it is possible that the NuoL-like and NuoM-like subunits are proton-translocating. It is likely that Hyf-like complexes can generate membrane potential through its antiporter-like subunits.
Sequences in this class
| NCBI Accession | Organism | Hydrogenase Class | Phylum | Order | Activity (Predicted) | Oxygen Tolerance (Predicted) | Subunits (Predicted) | Metal Centres (Predicted) | Accessory Subunits (Predicted) |
|---|---|---|---|---|---|---|---|---|---|
| WP_005366168.1 | Photobacterium angustum | [NiFe] Group 4a | Gammaproteobacteria | Vibrionales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_008989674.1 | Photobacterium leiognathi | [NiFe] Group 4a | Gammaproteobacteria | Vibrionales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_006232936.1 | Photobacterium profundum | [NiFe] Group 4a | Gammaproteobacteria | Vibrionales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_007469705.1 | Photobacterium sp. AK15 | [NiFe] Group 4a | Gammaproteobacteria | Vibrionales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_006644813.1 | Photobacterium sp. SKA34 | [NiFe] Group 4a | Gammaproteobacteria | Vibrionales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_004726128.1 | Vibrio furnissii | [NiFe] Group 4a | Gammaproteobacteria | Vibrionales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_021019734.1 | Vibrio gazogenes | [NiFe] Group 4a | Gammaproteobacteria | Vibrionales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_011751858.1 | Thermofilum pendens | [NiFe] Group 4a | Crenarchaeota | Thermoproteales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_022587809.1 | Caldanaerobacter subterraneus | [NiFe] Group 4a | Firmicutes | Thermoanaerobacterales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_025773400.1 | Moorella thermoacetica | [NiFe] Group 4a | Firmicutes | Thermoanaerobacterales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_015051661.1 | Thermacetogenium phaeum | [NiFe] Group 4a | Firmicutes | Thermoanaerobacterales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_029010114.1 | Azospirillum halopraeferens | [NiFe] Group 4a | Alphaproteobacteria | Rhodospirillales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_014249328.1 | Azospirillum lipoferum | [NiFe] Group 4a | Alphaproteobacteria | Rhodospirillales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_018609272.1 | Uliginosibacterium gangwonense | [NiFe] Group 4a | Betaproteobacteria | Rhodocyclales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_008384313.1 | Rhodovulum sp. PH10 | [NiFe] Group 4a | Alphaproteobacteria | Rhodobacterales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_011474970.1 | Rhodopseudomonas palustris | [NiFe] Group 4a | Alphaproteobacteria | Rhizobiales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_005542188.1 | Aggregatibacter actinomycetemcomitans | [NiFe] Group 4a | Gammaproteobacteria | Pasteurellales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_005703984.1 | Aggregatibacter aphrophilus | [NiFe] Group 4a | Gammaproteobacteria | Pasteurellales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_005645455.1 | Haemophilus haemolyticus | [NiFe] Group 4a | Gammaproteobacteria | Pasteurellales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_014065500.1 | Haemophilus parainfluenzae | [NiFe] Group 4a | Gammaproteobacteria | Pasteurellales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_007242420.1 | Haemophilus pittmaniae | [NiFe] Group 4a | Gammaproteobacteria | Pasteurellales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_005760524.1 | Pasteurella bettyae | [NiFe] Group 4a | Gammaproteobacteria | Pasteurellales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_005764546.1 | Pasteurella dagmatis | [NiFe] Group 4a | Gammaproteobacteria | Pasteurellales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_028536385.1 | Paludibacterium yongneupense | [NiFe] Group 4a | Betaproteobacteria | Neisseriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_007473606.1 | Caminibacter mediatlanticus | [NiFe] Group 4a | Epsilonproteobacteria | Nautiliales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_014558423.1 | Fervidicoccus fontis | [NiFe] Group 4a | Crenarchaeota | Fervidicoccales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_026823431.1 | Arsenophonus nasoniae | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_009113583.1 | Brenneria sp. EniD312 | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_029095340.1 | Budvicia aquatica | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_016538796.1 | Cedecea davisae | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_012134832.1 | Citrobacter koseri | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_012907207.1 | Citrobacter rodentium | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_006687169.1 | Citrobacter youngae | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_007725151.1 | Cronobacter dublinensis | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_007792874.1 | Cronobacter malonaticus | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_024559362.1 | Cronobacter pulveris | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_012124960.1 | Cronobacter sakazakii | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_015741042.1 | Cronobacter turicensis | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_013317233.1 | Dickeya dadantii | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_024105256.1 | Dickeya dianthicola | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_022632933.1 | Dickeya solani | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_019844844.1 | Dickeya zeae | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_024524460.1 | Edwardsiella hoshinae | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_015872494.1 | Edwardsiella ictaluri | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_005281999.1 | Edwardsiella tarda | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_015370028.1 | Enterobacter aerogenes | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_029739528.1 | Enterobacter asburiae | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_006178188.1 | Enterobacter cancerogenus | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_020883834.1 | Enterobacter cloacae | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
| WP_006811737.1 | Enterobacter hormaechei | [NiFe] Group 4a | Gammaproteobacteria | Enterobacteriales | Evolving | Sensitive | 6 or 9 | [NiFe]-centre, 1 x [4Fe4S] clusters | NuoL, NuoH, two [FeS] proteins, (three antiporter subunits) |
Literature
Genetics:
- Andrews, S.C., Berks, B.C., McClay, J., Ambler, A., Quail, M.A., Golby, P., and Guest, J.R. (1997) A 12-cistron Escherichia coli operon (hyf) encoding a putative proton-translocating formate hydrogenlyase system. Microbiol. 143: 3633-3647.
- Barrett, E.L., Kwan, H.S., and Macy, J. (1984) Anaerobiosis, formate, nitrate, and pyrA are involved in the regulation of formate hydrogenlyase in Salmonella typhimurium. J. Bacteriol. 158: 972-977.
- Böhm, R., Sauter, M., and Böck, A. (1990) Nucleotide sequence and expression of an operon in Escherichia coli coding for formate hydrogenylase components. Mol. Microbiol. 4: 231-243.
- Chippaux, M., Casse, F. and Pascal, M.C. (1972) Isolation and phenotypes of mutants from Salmonella typhimurium defective in formate hydrogenlyase activity. J. Bacteriol. 110: 766-768.
- Greening, C., Biswas, A., Carere, C.R., Jackson, C.J., Taylor, M.C., Stott, M.B., Cook, G.M., and Morales, S.E. (2016) Genomic and metagenomic surveys of hydrogenase distribution indicate H2 is a widely utilised energy source for microbial growth and survival. ISME J. 10: 761-777.
- Kruse, S., Goris, T., Wolf, M., Wei, X., and Diekert, G. (2017). The NiFe hydrogenases of the tetrachloroethene-respiring Epsilonproteobacterium Sulfurospirillum multivorans: biochemical studies and transcription analysis. Front. Microbiol. 8: 444.
- Marreiros, B.C., Batista, A.P., Duarte, A.M., and Pereira, M.M. (2013) A missing link between complex I and group 4 membrane-bound [NiFe] hydrogenases. BBA Bioenerg. 1827: 198-209.
- Pi, J., Jawed, M., Wang, J., Xu, L., and Yan, Y. (2016) Mutational analysis of the hyc-operon determining the relationship between hydrogenase-3 and NADH pathway in Enterobacter aerogenes. Enzyme Microb. Technol. 82: 1-7.
- Sauter, M., Böhm, R., and Böck, A. (1992) Mutational analysis of the operon (hyc) determining hydrogenase 3 formation in Escherichia coli. Mol. Microbiol. 6: 1523-1532.
- Sawers, R.G., Ballantine, S.P., and Boxer, D.H. (1985) Differential expression of hydrogenase isoenzymes in Escherichia coli K-12: evidence for a third isoenzyme. J. Bacteriol. 164: 1324-1331.
- Schlensog, V. and Böck, A. (1990) Identification and sequence analysis of the gene encoding the transcriptional activator of the formate hydrogenlyase system of Escherichia coli. Mol. Microbiol. 4: 1319-1327.
- Self, W.T., Hasona, A. and Shanmugam, K.T. (2004) Expression and regulation of a silent operon, hyf, coding for hydrogenase 4 isoenzyme in Escherichia coli. J. Bacteriol. 186: 580-587.
- Skibinski, D.A., Golby, P., Chang, Y.S., Sargent, F., Hoffman, R., Harper, R., Guest, J.R., Attwood, M.M., Berks, B.C., and Andrews, S.C. (2002) Regulation of the hydrogenase-4 operon of Escherichia coli by the sigma54-dependent transcriptional activators FhlA and HyfR. J. Bacteriol. 184: 6642-6653.
- Zinoni, F., Birkmann, A., Stadtman, T.C., and Böck, A. (1986) Nucleotide sequence and expression of the selenocysteine-containing polypeptide of formate dehydrogenase (formate-hydrogen-lyase-linked) from Escherichia coli. Proc. Acad. Natl. Sci. USA 83: 4650-4654.
Physiology:
- Lamichhane-Khadka, R., Benoit, S.L., Miller-Parks, E.F., and Maier, R.J. (2015) Host hydrogen rather than that produced by the pathogen is important for Salmonella enterica serovar Typhimurium virulence. Infect. Immun. 83: 311-316.
- Mnatsakanyan, N., Bagramyan, K., and Trchounian, A. (2004) Hydrogenase 3 but not hydrogenase 4 is major in hydrogen gas production by Escherichia coli formate hydrogenlyase at acidic pH and in the presence of external formate. Cell Biochem. Biophys. 41: 357-365.
- Noguchi, K., Riggins, D.P., Eldahan, K.C., Kitko, R.D., and Slonczewski, J.L. (2010) Hydrogenase-3 contributes to anaerobic acid resistance of Escherichia coli. PLOS One 5: e10132.
- Peck Jr, H.D. and Gest, H. (1957) Formic dehydrogenase and the hydrogenlyase enzyme complex in coli-aerogenes bacteria. J. Bacteriol. 73: 706-721.
- Redwood, M.D., Mikheenko, I.P., Sargent, F. and Macaskie, L.E. (2008) Dissecting the roles of Escherichia coli hydrogenases in biohydrogen production. FEMS Microbiol. Lett. 278: 48-55.
- Qadri, S.H. and Hoare, D.S. (1968) Formic hydrogenlyase and the photoassimilation of formate by a strain of Rhodopseudomonas palustris. J. Bacteriol. 95: 2344-2357.
- Roger, M., Brown, F., Gabrielli, W., and Sargent, F. (2018) Efficient hydrogen-dependent carbon dioxide reduction by Escherichia coli. Curr. Biol. 8: 140-145.
- Sawers, R.G., Jamieson, D.J., Higgins, C.F., and Boxer, D.H. (1986) Characterization and physiological roles of membrane-bound hydrogenase isoenzymes from Salmonella typhimurium. J. Bacteriol. 168: 398-404.
- Stephenson, M. and Stickland, L.H. (1932) Hydrogenlyases: Bacterial enzymes liberating molecular hydrogen. Biochemical J. 26: 712-724.
- Trchounian, K. and Trchounian, A. (2013) Escherichia coli hydrogenase 4 (hyf) and hydrogenase 2 (hyb) contribution in H2 production during mixed carbon (glucose and glycerol) fermentation at pH 7.5 and pH 5.5. Int. J. Hydrogen Energy 38: 3921-3929.
- Trchounian, K. and Trchounian, A. (2014) Hydrogen producing activity by Escherichia coli hydrogenase 4 (hyf) depends on glucose concentration. Int. J. Hydrogen Energy 39: 16914-16918.
- Yoshida, A., Nishimura, T., Kawaguchi, H., Inui, M., and Yukawa, H. (2005) Enhanced hydrogen production from formic acid by formate hydrogen lyase-overexpressing Escherichia coli strains. Appl. Environ. Microbiol. 71: 6762-6768.
Biochemistry:
- Axley, M.J., Grahame, D.A., and Stadtman, T.C. (1990) Escherichia coli formate-hydrogen lyase. Purification and properties of the selenium-dependent formate dehydrogenase component. J. Biol. Chem. 265: 18213-18218.
- Maeda, T., Sanchez-Torres, V., and Wood, T.K. (2007) Escherichia coli hydrogenase 3 is a reversible enzyme possessing hydrogen uptake and synthesis activities. Appl. Microbiol. Biotechnol. 76: 1035-1042.
- McDowall, J.S., Hjersing, M.C., Palmer, T., and Sargent, F. (2015) Dissection and engineering of the Escherichia coli formate hydrogenlyase complex. FEBS Lett. 589: 3141-3147.
- McDowall, J.S., Murphy, B.J., Haumann, M., Palmer, T., Armstrong, F.A., and Sargent, F. (2014) Bacterial formate hydrogenlyase complex. Proc. Natl. Acad. Sci. USA 111: E3948-E3956.
- Lamont, C.M., Kelly, C.L., Pinske, C., Buchanan, G., Palmer, T., and Sargent, F. (2017) Expanding the substrates for a bacterial hydrogenlyase reaction. Microbiol. 163: 649-653.
- Pinske, C. and Sargent, F. (2016) Exploring the directionality of Escherichia coli formate hydrogenlyase: a membrane-bound enzyme capable of fixing carbon dioxide to organic acid. MicrobiologyOpen 5: 721-737.