[NiFe] Group 3d-hydrogenase
This entry was last updated at: Feb. 9, 2018, 6:37 a.m.
Properties
| Group | [NiFe] Group 3: Cofactor-coupled bidirectional [NiFe] hydrogenases |
| Subgroup | [NiFe] Group 3d: NAD-coupled |
| Function | Generates reductant for hydrogenotrophic carbon-fixation by directly coupling oxidation of H2 to reduction of NAD. Also acts in reverse direction to catalyse fermentative NADH-dependent production of H2. Electrons flow between H2-activating hydrogenase subunits to NAD-activating diaphorase subunits. |
| Activity | Bidirectional |
| Oxygen tolerance | O2-tolerant |
| Localisation | Cytosolic |
Distribution
| Ecosystem distribution | Diverse soil and aquatic environments |
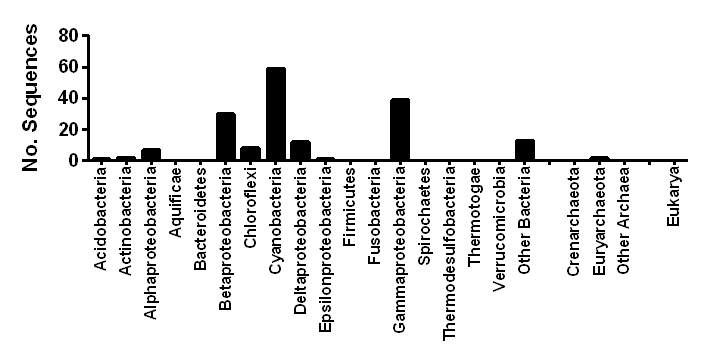
| Taxonomic distribution | Encoded in diverse Proteobacteria and Cyanobacteria |
Genetic Organisation
e.g. Ralstonia eutropha (WP_011154013.1):

Architecture
| Structures | |
| Subunits | 4 or 5 |
| Subunit description | HoxH (hydrogenase large subunit) HoxY (hydrogenase small subunit) HoxF (diaphorase catalytic subunit) HoxU (diaphorase electron transfer subunit) Photosynthetic bacteria also contain HoxE (iron-sulfur subunit) |
| Catalytic site | [NiFe]-centre |
| FeS clusters | Proximal: 4Cys[4Fe4S] Medial: Absent Distal: Absent |
Sequences in this class
| NCBI Accession | Organism | Hydrogenase Class | Phylum | Order | Activity (Predicted) | Oxygen Tolerance (Predicted) | Subunits (Predicted) | Metal Centres (Predicted) | Accessory Subunits (Predicted) |
|---|---|---|---|---|---|---|---|---|---|
| WP_017839865.1 | Methylomicrobium buryatense | [NiFe] Group 3d | Gammaproteobacteria | Methylococcales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_013816969.1 | Methylomonas methanica | [NiFe] Group 3d | Gammaproteobacteria | Methylococcales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_026601561.1 | Methylomonas sp. 11b | [NiFe] Group 3d | Gammaproteobacteria | Methylococcales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_020482704.1 | Methylomonas sp. MK1 | [NiFe] Group 3d | Gammaproteobacteria | Methylococcales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_020565381.1 | Methylosarcina fibrata | [NiFe] Group 3d | Gammaproteobacteria | Methylococcales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_024299641.1 | Methylosarcina lacus | [NiFe] Group 3d | Gammaproteobacteria | Methylococcales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_019865758.1 | Methylovulum miyakonense | [NiFe] Group 3d | Gammaproteobacteria | Methylococcales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_011394103.1 | Hahella chejuensis | [NiFe] Group 3d | Gammaproteobacteria | Oceanospirillales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_020410523.1 | Hahella ganghwensis | [NiFe] Group 3d | Gammaproteobacteria | Oceanospirillales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_020161995.1 | Cycloclasticus pugetii | [NiFe] Group 3d | Gammaproteobacteria | Thiotrichales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_015005599.1 | Cycloclasticus sp. P1 | [NiFe] Group 3d | Gammaproteobacteria | Thiotrichales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_016389591.1 | Cycloclasticus sp. PY97M | [NiFe] Group 3d | Gammaproteobacteria | Thiotrichales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_020932782.1 | Cycloclasticus zancles | [NiFe] Group 3d | Gammaproteobacteria | Thiotrichales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_020393771.1 | Thiothrix disciformis | [NiFe] Group 3d | Gammaproteobacteria | Thiotrichales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_029133296.1 | Sedimenticola selenatireducens | [NiFe] Group 3d | Gammaproteobacteria | Unclassified | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_005505390.1 | Grimontia hollisae | [NiFe] Group 3d | Gammaproteobacteria | Vibrionales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_029165455.1 | Aminiphilus circumscriptus | [NiFe] Group 3d | Synergistetes | Synergistales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_013048933.1 | Aminobacterium colombiense | [NiFe] Group 3d | Synergistetes | Synergistales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_024821667.1 | Aminobacterium mobile | [NiFe] Group 3d | Synergistetes | Synergistales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_009200281.1 | Anaerobaculum hydrogeniformans | [NiFe] Group 3d | Synergistetes | Synergistales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_014807493.1 | Anaerobaculum mobile | [NiFe] Group 3d | Synergistetes | Synergistales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_005661842.1 | Dethiosulfovibrio peptidovorans | [NiFe] Group 3d | Synergistetes | Synergistales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_014163745.1 | Thermovirga lienii | [NiFe] Group 3d | Synergistetes | Synergistales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
| WP_009849435.1 | Mariprofundus ferrooxydans | [NiFe] Group 3d | Zetaproteobacteria | Mariprofundales | Bidirectional | Tolerant | 4 | [NiFe]-centre, 1 x [4Fe4S] clusters | Diaphorase, [FeS] protein |
Literature
Genetics:
- Boison, G., Bothe, H., and Schmitz, O. (2000) Transcriptional analysis of hydrogenase genes in the cyanobacteria Anacystis nidulans and Anabaena variabilis monitored by RT-PCR. Curr. Microbiol. 40: 315-321.
- Boison, G., Schmitz, O., Mikheeva, L., Shestakov, S., and Bothe, H. (1996) Cloning, molecular analysis and insertional mutagenesis of the bidirectional hydrogenase genes from the cyanobacterium Anacystis nidulans. FEBS Lett. 394: 153-158.
- Boison, G., Schmitz, O., Schmitz, B., and Bothe, H. (1998) Unusual gene arrangement of the bidirectional hydrogenase and functional analysis of its diaphorase subunit HoxU in respiration of the unicellular cyanobacterium Anacystis nidulans. Curr. Microbiol. 36: 253-258.
- Eberz, G., Hogrefe, C., Kortlüke, C., Kamienski, A., and Friedrich, B. (1986) Molecular cloning of structural and regulatory hydrogenase (hox) genes of Alcaligenes eutrophus H16. J. Bacteriol. 168: 636-641.
- Eckert, C., Boehm, M., Carrieri, D., Yu, J., Dubini, A., Nixon, P.J., and Maness, P.C. (2012) Genetic analysis of the Hox hydrogenase in the cyanobacterium Synechocystis sp. PCC 6803 reveals subunit roles in association, assembly, maturation, and function. J. Biol. Chem. 287: 43502-43515.
- Gutekunst, K., Phunpruch, S., Schwarz, C., Schuchardt, S., Schulz-Friedrich, R., and Appel, J. (2005) LexA regulates the bidirectional hydrogenase in the cyanobacterium Synechocystis sp. PCC 6803 as a transcription activator. Mol. Microbiol. 58: 810-823.
- Grzeszik, C., Lübbers, M., Reh, M. and Schlegel, H.G. (1997) Genes encoding the NAD-reducing hydrogenase of Rhodococcus opacus MR11. Microbiol. 143: 1271-1286.
- Hogrefe, C., Römermann, D., and Friedrich, B. (1984) Alcaligenes eutrophus hydrogenase genes (Hox). J. Bacteriol. 158: 43-48.
- Maróti, J., Farkas, A., Nagy, I.K., Maróti, G., Kondorosi, É., Rákhely, G., and Kovács, K.L. (2010) A second soluble Hox-type NiFe enzyme completes the hydrogenase set in Thiocapsa roseopersicina BBS. Appl. Environ. Microbiol. 76: 5113-5123.
- Oliveira, P. and Lindblad, P. (2005) LexA, a transcription regulator binding in the promoter region of the bidirectional hydrogenase in the cyanobacterium Synechocystis sp. PCC 6803. FEMS Microbiol. Lett. 251: 59-66.
- Oliveira, P. and Lindblad, P. (2009) Transcriptional regulation of the cyanobacterial bidirectional Hox-hydrogenase. Dalton Trans. 45: 9990-9996.
- Schmitz, O., Boison, G., and Bothe, H. (2001) Quantitative analysis of expression of two circadian clock-controlled gene clusters coding for the bidirectional hydrogenase in the cyanobacterium Synechococcus sp. PCC7942. Mol. Microbiol. 41: 1409-1417.
- Schmitz, O., Boison, G., Hilscher, R., Hundeshagen, B., Zimmer, W., Lottspeich, F., and Bothe, H. (1995) Molecular biological analysis of a bidirectional hydrogenase from cyanobacteria. Eur. J. Biochem. 233: 266-276.
- Sheremetieva, M.E., Troshina, O.Y., Serebryakova, L.T., and Lindblad, P. (2002) Identification of hox genes and analysis of their transcription in the unicellular cyanobacterium Gloeocapsa alpicola CALU 743 growing under nitrate-limiting conditions. FEMS Microbiol. Lett. 214: 229-233
- Sjöholm, J., Oliveira, P., and Lindblad, P. (2007) Transcription and regulation of the bidirectional hydrogenase in the cyanobacterium Nostoc sp. strain PCC 7120. Appl. Environ. Microbiol. 73: 5435-5446
- Tran-Betcke, A., Warnecke, U., Böcker, C., Zaborosch, C., and Friedrich, B. (1990) Cloning and nucleotide sequences of the genes for the subunits of NAD-reducing hydrogenase of Alcaligenes eutrophus H16. J. Bacteriol. 172: 2920-2929.
- Vignais, P.M., Billoud, B., and Meyer, J. (2001) Classification and phylogeny of hydrogenases. FEMS Microbiol. Rev. 25: 455-501.
- Wells, M.A., Mercer, J., Mott, R.A., Pereira-Medrano, A.G., Burja, A.M., Radianingtyas, H., and Wright, P.C. (2011) Engineering a non-native hydrogen production pathway into Escherichia coli via a cyanobacterial [NiFe] hydrogenase. Metab. Eng. 13: 445-453.
Physiology:
- Aggag, M. and Schlegel, H.G. (1973) Studies on a gram-positive hydrogen bacterium, Nocardia opaca strain 1b. Arch. Microbiol. 88: 299-318.
- Antal, T.K. and Lindblad, P. (2005) Production of H2 by sulphur-deprived cells of the unicellular cyanobacteria Gloeocapsa alpicola and Synechocystis sp. PCC 6803 during dark incubation with methane or at various extracellular pH. J. Appl. Microbiol. 98: 114-120.
- Aubert-Jousset, E., Cano, M., Guedeney, G., Richaud, P., and Cournac, L. (2011) Role of HoxE subunit in Synechocystis PCC6803 hydrogenase. FEBS J. 278: 4035-4043.
- Boison, G., Bothe, H., Hansel, A., and Lindblad, P. (1999) Evidence against a common use of the diaphorase subunits by the bidirectional hydrogenase and by the respiratory complex I in cyanobacteria. FEMS Microbiol. Lett. 174: 159-165.
- Carrieri, D., Wawrousek, K., Eckert, C., Yu, J., and Maness, P.C. (2011) The role of the bidirectional hydrogenase in cyanobacteria. Bioresource Technol. 102: 8368-8377.
- De Rosa, E., Checchetto, V., Franchin, C., Bergantino, E., Berto, P., Szabò, I., Giacometti, G.M., Arrigoni, G., and Costantini, P. (2015) [NiFe]-hydrogenase is essential for cyanobacterium Synechocystis sp. PCC 6803 aerobic growth in the dark. Sci. Rep. 5: 12424.
- Friedrich, C.G., Friedrich, B. and Bowien, B. (1981) Formation of enzymes of autotrophic metabolism during heterotrophic growth of Alcaligenes eutrophus. Microbiol. 122: 69-78.
- Grzeszik, C., Ross, K., Schneider, K., Reh, M., and Schlegel, H.G. (1997) Location, catalytic activity, and subunit composition of NAD-reducing hydrogenases of some Alcaligenes strains and Rhodococcus opacus MR22. Arch. Microbiol. 167: 172-176.
- Gutekunst, K., Chen, X., Schreiber, K., Kaspar, U., Makam, S., and Appel, J. (2014) The bidirectional NiFe-hydrogenase in Synechocystis sp. PCC 6803 is reduced by flavodoxin and ferredoxin and is essential under mixotrophic, nitrate-limiting conditions. J. Biol. Chem. 289: 1930-1937.
- Hanczár, T., Csáki, R., Bodrossy, L., Murrell, C.J., and Kovács, K.L. (2002) Detection and localization of two hydrogenases in Methylococcus capsulatus (Bath) and their potential role in methane metabolism. Arch. Microbiol. 177: 167-172.
- Houchins, J.P. and Burris, R.H. (1981) Occurrence and localization of two distinct hydrogenases in the heterocystous cyanobacterium Anabaena sp. strain 7120. J. Bacteriol. 146: 209-214.
- Laurinavichene, T.V., Rákhley, G., Kovács, K.L., and Tsygankov, A.A. (2007) The effect of sulfur compounds on H2 evolution/consumption reactions, mediated by various hydrogenases, in the purple sulfur bacterium, Thiocapsa roseopersicina. Arch. Microbiol. 188: 403-410.
- Mori, J.F., Scott, J.J., Hager, K.W., Moyer, C.L., Küsel, K., and Emerson, D. (2017) Physiological and ecological implications of an iron-or hydrogen-oxidizing member of the Zetaproteobacteria, Ghiorsea bivora, gen. nov., sp. nov. ISME J. 11: 2624-2636.
- Pálágyi-Mészáros, L.S., Maróti, J., Latinovics, D., Balogh, T., Klement, E., Medzihradszky, K.F., Rákhely, G., and Kovács, K.L. (2009) Electron-transfer subunits of the NiFe hydrogenases in Thiocapsa roseopersicina BBS. FEBS J. 276: 164-174.
- Papen, H., Kentemich, T., Schmülling, T. and Bothe, H. (1986) Hydrogenase activities in cyanobacteria. Biochimie 68: 121-132.
- Porthun, A., Bernhard, M., and Friedrich, B. (2002) Expression of a functional NAD-reducing [NiFe] hydrogenase from the gram-positive Rhodococcus opacus in the gram-negative Ralstonia eutropha. Arch. Microbiol. 177: 159-166.
- Rákhely, G., Kovács, A.T., Maroti, G., Fodor, B.D., Csanádi, G., Latinovics, D., and Kovács, K.L. (2004) Cyanobacterial-type, heteropentameric, NAD+-reducing NiFe hydrogenase in the purple sulfur photosynthetic bacterium Thiocapsa roseopersicina. Appl. Environ. Microbiol. 70: 722-728.
- Rákhely, G., Laurinavichene, T.V., Tsygankov, A.A., and Kovács, K.L. (2007) The role of Hox hydrogenase in the H2 metabolism of Thiocapsa roseopersicina. BBA Bioenerg. 1767: 671-676.
- Schmitz, O. and Bothe, H. (1996) NAD(P)+-dependent hydrogenase activity in extracts from the cyanobacterium Anacystis nidulans. FEMS Microbiol. Lett. 135: 97-101.
- Schütz, K., Happe, T., Troshina, O., Lindblad, P., Leitão, E., Oliveira, P., and Tamagnini, P. (2004) Cyanobacterial H2 production—a comparative analysis. Planta 218: 350-359.
- Tamagnini, P., Leitão, E., Oliveira, P., Ferreira, D., Pinto, F., Harris, D.J., Heidorn, T., and Lindblad, P. (2007) Cyanobacterial hydrogenases: diversity, regulation and applications. FEMS Microbiol. Rev. 31: 692-720.
Biochemistry:
- Burgdorf, T., van der Linden, E., Bernhard, M., Yin, Q.Y., Back, J.W., Hartog, A.F., Muijsers, A.O., de Koster, C.G., Albracht, S.P., and Friedrich, B. (2005) The soluble NAD+-reducing [NiFe]-hydrogenase from Ralstonia eutropha H16 consists of six subunits and can be specifically activated by NADPH. J. Bacteriol. 187: 3122-3132.
- Germer, F., Zebger, I., Saggu, M., Lendzian, F., Schulz, R., and Appel, J. (2009) Overexpression, isolation, and spectroscopic characterization of the bidirectional [NiFe] hydrogenase from Synechocystis sp. PCC 6803. J. Biol. Chem. 284: 36462-36472.
- Karstens, K., Wahlefeld, S., Horch, M., Grunzel, M., Lauterbach, L., Lendzian, F., Zebger, I., and Lenz, O. (2015) Impact of the iron-sulfur cluster proximal to the active site on the catalytic function of an O2-tolerant NAD+-reducing [NiFe]-hydrogenase. Biochem. 54: 389-403.
- Horch, M., Lauterbach, L., Lenz, O., Hildebrandt, P., and Zebger, I. (2012) NAD(H)-coupled hydrogen cycling: structure-function relationships of bidirectional [NiFe] hydrogenases. FEBS Lett. 586: 545-556.
- Horch, M., Lauterbach, L., Mroginski, M.A., Hildebrandt, P., Lenz, O., and Zebger, I. (2015) Reversible active site sulfoxygenation can explain the oxygen tolerance of a NAD+-reducing [NiFe] hydrogenase and its unusual infrared spectroscopic properties. J. Am. Chem. Soc. 137: 2555-2564.
- Lauterbach, L. and Lenz, O. (2013) Catalytic production of hydrogen peroxide and water by oxygen-tolerant [NiFe]-hydrogenase during H2 cycling in the presence of O2. J. Am. Chem. Soc. 135: 17897-17905.
- Lauterbach, L., Idris, Z., Vincent, K.A., and Lenz, O. (2011) Catalytic properties of the isolated diaphorase fragment of the NAD+-reducing [NiFe]-hydrogenase from Ralstonia eutropha. PLOS One 6: e25939.
- Lauterbach, L., Lenz, O. and Vincent, K.A. (2013) H2-driven cofactor regeneration with NAD(P)+-reducing hydrogenases. FEBS J. 280: 3058-3068.
- Lauterbach, L., Liu, J., Horch, M., Hummel, P., Schwarze, A., Haumann, M., Vincent, K.A., Lenz, O., and Zebger, I. (2011) The hydrogenase subcomplex of the NAD+-reducing [NiFe] hydrogenase from Ralstonia eutropha—insights into catalysis and redox interconversions. Eur. J. Inorg. Chem. 7: 1067-1079.
- Löscher, S., Burgdorf, T., Zebger, I., Hildebrandt, P., Dau, H., Friedrich, B., and Haumann, M. (2006) Bias from H2 cleavage to production and coordination changes at the Ni-Fe active site in the NAD+-reducing hydrogenase from Ralstonia eutropha. Biochem. 45: 11658-11665.
- Massanz, C. and Friedrich, B. (1999) Amino acid replacements at the H2-activating site of the NAD-reducing hydrogenase from Alcaligenes eutrophus. Biochem. 38: 14330-14337.
- McIntosh, C.L., Germer, F., Schulz, R., Appel, J., and Jones, A.K. (2011) The [NiFe]-hydrogenase of the cyanobacterium Synechocystis sp. PCC 6803 works bidirectionally with a bias to H2 production. J. Am. Chem. Soc. 133: 11308-11319.
- Preissler, J., Wahlefeld, S., Lorent, C., Teutloff, C., Horch, M., Lauterbach, L., Cramer, S.P., Zebger, I., and Lenz, O. (2018) Enzymatic and spectroscopic properties of a thermostable [NiFe]‑hydrogenase performing H2-driven NAD+-reduction in the presence of O2. BBA Bioenerg. 1859: 8-18.
- Ratzka, J., Lauterbach, L., Lenz, O., and Ansorge-Schumacher, M.B. (2011) Systematic evaluation of the dihydrogen-oxidising and NAD+-reducing soluble [NiFe]-hydrogenase from Ralstonia eutropha H16 as a cofactor regeneration catalyst. Biocat. Biotransf. 29: 246-252.
- Schmitz, O., Boison, G., Salzmann, H., Bothe, H., Schütz, K., Wang, S.H., and Happe, T (2002) HoxE - a subunit specific for the pentameric bidirectional hydrogenase complex (HoxEFUYH) of cyanobacteria. BBA Bioenerg. 1554: 66-74.
- Schneider, K. and Schlegel, H.G. (1976) Purification and properties of soluble hydrogenase from Alcaligenes eutrophus H16. BBA Enzymol. 452: 66-80.
- Schneider, K., Cammack, R., and Schlegel, H.G. (1984) Content and localization of FMN, Fe-S clusters and nickel in the NAD-linked hydrogenase of Nocardia opaca 1b. Eur. J. Biochem. 142: 75-84.
- Schneider, K., Cammack, R., Schlegel, H.G., and Hall, D.O. (1979) The iron-sulphur centres of soluble hydrogenase from Alcaligenes eutrophus. BBA Prot. Struct. 578: 445-461.
- Shomura, Y., Taketa, M., Nakashima, H., Tai, H., Nakagawa, H., Ikeda, Y., Ishii, M., Igarashi, Y., Nishihara, H., Yoon, K.S., Ogo, S., Hirota, S., and Higuchi, Y. (2017) Structural basis of the redox switches in the NAD+-reducing soluble [NiFe]-hydrogenase. Science 357: 928-932
Chemistry:
- Erkens, A., Schneider, K., and Müller, A. (1996) The NAD-linked soluble hydrogenase from Alcaligenes eutrophus H16: detection and characterization of EPR signals deriving from nickel and flavin. J. Biol. Inorg. Chem. 1: 99-110.
- Long, M., Liu, J., Chen, Z., Bleijlevens, B., Roseboom, W. and Albracht, S.P. (2007) Characterization of a HoxEFUYH type of [NiFe] hydrogenase from Allochromatium vinosum and some EPR and IR properties of the hydrogenase module. J. Biol. Inorg. Chem. 12: 62-78.
- Horch, M., Lauterbach, L., Saggu, M., Hildebrandt, P., Lendzian, F., Bittl, R., Lenz, O., and Zebger, I., 2010. Probing the active site of an O2-tolerant NAD+-reducing [NiFe]-hydrogenase from Ralstonia eutropha H16 by in situ EPR and FTIR spectroscopy. Angewandte Chemie 49: 8026-8029.
- Horch, M., Rippers, Y., Mroginski, M.A., Hildebrandt, P., and Zebger, I. (2013) Combining spectroscopy and theory to evaluate structural models of metalloenzymes: a case study on the soluble [NiFe] hydrogenase from Ralstonia eutropha. ChemPhysChem 14: 185-191.
- Lauterbach, L., Wang, H., Horch, M., Gee, L.B., Yoda, Y., Tanaka, Y., Zebger, I., Lenz, O., and Cramer, S.P. (2015) Nuclear resonance vibrational spectroscopy reveals the FeS cluster composition and active site vibrational properties of an O2-tolerant NAD+-reducing [NiFe] hydrogenase. Chem. Sci. 6: 1055-1060.
- Löwenstein, J., Lauterbach, L., Teutloff, C., Lenz, O., and Bittl, R. (2015). Active site of the NAD+-reducing hydrogenase from Ralstonia eutropha studied by EPR spectroscopy. J. Phys. Chem. B 119: 13834-13841.